FEF-IA Ghana : Artificial Intelligence For Sustainable Development – Institut de Chimie Physique, Université Paris Saclay/ CNRS UMR8000
Thematic: Act 1.1: Integrative biology, (bio)chemistry, and biophysics to support research and human resource development
The Institut de Chimie Physique (ICP) of Université Paris-Saclay has recently initiated its activities within the framework of the FEF-IA Ghana: Artificial Intelligence for Sustainable Development in Ghana project. The initial phase of the project has welcomed three students from Ghana, one Ghanaian student already based in France, and an Ingénieur d’Étude recruited under the project.
Additional students from Ghana are expected to join the French team in France in the coming months to contribute to the execution of the project’s thematic areas. These activities are primarily focused on the application of artificial intelligence to health-related research, particularly in the fields of biophysics and bio(chemistry).
Within the FEF-IA project, students and researchers are assigned specific tasks aligned with the overall objectives of the initiative. These tasks are designed to promote skill development, interdisciplinary collaboration, and the practical application of artificial intelligence in scientific research.
The following sections describe the mini-projects currently being investigated and developed by each student.

1. Sandra Agbenyegah
“Comparative Study of Conventional and FLASH Radiotherapy on inflammatory signalling, and downstream cellular responses: Improving data exploitation and analysis using AI.”
Sandra Agbenyegah is a PhD student in Health Physics at the University of Ghana and a Principal Technologist at the Applied Radiation Biology Center (ARBC) of the Radiological and Medical Sciences Research Institute (RAMSRI), Ghana Atomic Energy Commission (GAEC). GAEC is one of the partners of the FEF-IA.
Under the FEF-IA project, Sandra is working on the research topic entitled: “Comparative Study of Conventional and FLASH Radiotherapy on inflammatory signalling, and downstream cellular responses: Improving data exploitation and analysis using AI.”
The primary objective of her study is to experimentally compare the effects of conventional radiotherapy and FLASH radiotherapy, particularly with respect to immune defences. It is well known that immune defences are the major problem encountered after radiotherapy. The evaluation of dose and dose rate, on NADPH oxidase deactivation and reactive oxygen species (ROS) generation in neutrophil membranes has to be better known before being used for the development and evaluation of AI-based predictive models, including machine learning classifiers and neural networks, aimed at predicting functional alteration or preservation of phagocytes following exposure to conventional or FLASH radiotherapy.
Sandra arrived at the Institut de Chimie Physique (ICP), Université Paris-Saclay, on 1 November 2025. Since her arrival, she has undertaken several experimental activities and begun to be trained in the use of ionising radiations. These include Fricke dosimetry, a well-established chemical dosimetry technique used to measure absorbed radiation dose and dose-rate based on radiation-induced chemical changes in an aqueous solution. This method relies on the oxidation of ferrous ions (Fe²⁺) to ferric ions (Fe³⁺) upon exposure to ionizing radiation.

a. Lipid Extraction

b. Lipid Layer
This on-going mass spectrometry–based lipidomics at the Laboratoire de Chimie Moléculaire (LCM), École Polytechnique, in collaboration with Dr. Touboul led to large amount of data aiming at the identification and quantification of specific lipids induced under several different treatment conditions. Analyses aim to identify treatment-specific lipid signatures. The use of high-resolution mass spectrometry enabled comprehensive lipid characterization, including the detection of lipid species associated with lipid peroxidation and oxidative stress.
Future Work: With the experimental protocols now finalized, the next phase of the project will build upon the preliminary work through systematic experimentation and data acquisition, with particular emphasis on ensuring reproducibility and biological relevance of the results. Pilot experiments have already commenced to validate the dosimetry protocols and to assess experimental variability under both conventional and FLASH radiotherapy conditions, particularly in relation to neutrophil response and lipid peroxidation.
In parallel, further optimization of laboratory methodologies will be undertaken. This includes refinement of lipid extraction procedures, optimization of mass spectrometry parameters, and strengthening of quality control measures to ensure the generation of consistent and high-quality lipidomic data. Following the completion of this phase, large-scale data collection will be performed to directly compare the effects of conventional and FLASH radiotherapy on NADPH oxidase activity and ROS production in neutrophils. The resulting datasets will form the foundation for comprehensive statistical analysis, biological interpretation, and the development of AI-driven predictive models.
2. Miss Amanda Danumbu and Mr Rexford Tutu

Miss Amanda Danumbu

Mr. Rexford Tutu
Miss Amanda Danumbu and Mr. Rexford Tutu were selected under the FEF-IA project to work on the research topics entitled “Modelling Gold Nanoparticle–Water Interface under Ionizing Radiation” and “Modelling Nucleosomal DNA Irradiation by Fast Ions,” respectively.
Both Amanda and Rexford are Teaching Assistants at Kwame Nkrumah University of Science and Technology (KNUST). Amanda recently completed her Bachelor of Science degree in Chemistry, with a specialization in Computational Chemistry and Machine Learning. Mr. Rexford Tutu completed his Bachelor of Science degree in Physics, majoring in Biomedical Physics, graduating with First Class Honours. He has been actively involved in the application of machine learning techniques in medical physics.
Amanda and Rexford were recommended by their respective departments for an internship under the FEF-IA project, based on their academic excellence and strong interest in computational modelling and artificial intelligence. They arrived at the Institut de Chimie Physique (ICP), France, on 8 November 2025, and have since been actively engaged in their research activities.
Since their arrival, both students have been working on the development of AI models based on neural networks to predict the stopping powers of fast ions interacting with water in different physicochemical states, using data generated from first-principles simulations. This modelling effort is central to understanding radiation–matter interactions at the molecular and nanoscale levels, with direct relevance to radiobiology and medical physics applications.
The students are jointly supervised by Dr. Carine Clavaguera and Dr. Aurélien De La Lande, both are Directors of Research of CNRS. Over the past few months of their internship, Amanda and Rexford have intensively worked on programming environment for validation approach of using neural networks in predicting the stopping powers of fast ions in water. This included the installation and configuration of all necessary software packages, libraries, and dependencies needed to implement, train, and validate the neural network models.
The developed neural network model has been successfully tested and has demonstrated a potential ability to accurately predict stopping powers in water, as illustrated in Figure 1. This initial success provides a strong foundation for further model refinement and extension to more complex biological and nanostructured systems.
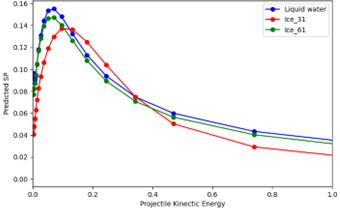
The current modelling framework will be further extended using real-time time-dependent density functional theory (RT-TDDFT) reference data for the training of advanced machine learning models. This approach will enable the prediction of electronic properties at large spatial and temporal scales, as well as the accurate estimation of stopping powers in complex biological systems, including DNA, proteins, and solvent environments.

A visit to the ICP cluster center
Amanda Danumbu and Rexford Tutu will return to Ghana on 31 January 2026, marking the completion of their three-month internship under the FEF-IA project. To ensure continuity of their research activities after their return, a high-performance computing (HPC) system has been purchased and installed at the Institut de Chimie Physique (ICP). This infrastructure has been configured to support remote access, thereby enabling all partner institutions to collaborate effectively and continue computational work at a distance. This arrangement will facilitate sustained scientific interaction, efficient data sharing, and continued development of AI-driven models within the framework of the FEF-IA project.
Amanda Danumbu and Rexford Tutu are expected to come back in May 2026 for another 3 month-internship to acquire knowledge in DFT and to continue the project on complex biological systems, including DNA, proteins, and solvent environments.
3. Octavio Bravo

Mr. Octavio Bravo was recruited under the FEF-IA project for the period from 1 November 2025 to 30 April 2026 an an Igénieur d’Etude. His research focuses on “AI-guided structural modelling of the NOX2 NADPH oxidase complex.”
Octavio’s work makes extensive use of AlphaFold3 (AF3), a state-of-the-art generative artificial intelligence model capable of predicting three-dimensional structures of biomolecules and multicomponent complexes directly from sequence information. His primary objectives are to (i) construct reliable structural models of protein assemblies using generative AI approaches, including AlphaFold2 and AlphaFold3, (ii) investigate structural rearrangements associated with activation of the NOX2 complex, an essential enzyme of the immune system, and (iii) develop in silico analysis tools to interrogate these structures, assess model stability, and establish meaningful connections between computational predictions and experimental data. Ultimately, this integrated approach aims to optimize experimental conditions and protein constructs.
To achieve these objectives, Octavio has initiated the development of custom Python programs for protein assembly analysis.
These tools are designed to quantify conserved inter-protein contacts and to group predicted structures into clusters based on per-chain root-mean-square deviation (RMSD) values. The analysis pipeline supports the evaluation of multiple prediction sets generated using different random seeds and enables systematic comparisons between models produced with and without structural templates.
For structural consistency, predicted assemblies are aligned on the transmembrane module of the NOX2 complex, allowing the specific assessment of conformational variability within the cytosolic regions. RMSD values are calculated for the transmembrane components and, independently, for the p47, p67 and Rac subunits, providing a detailed and quantitative framework for comparing structural dynamics across model ensembles.
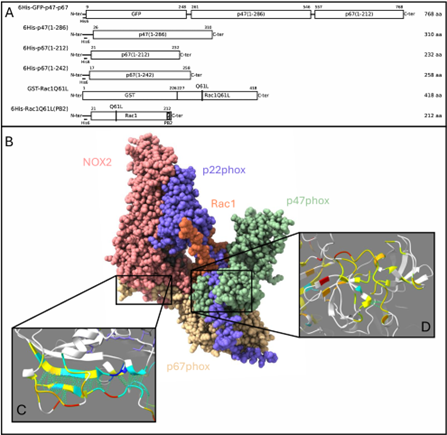
Figure 2. (A) Schematic representations of the recombinant cytosolic constructs used for functional assays. GST, glutathione S-transferase. GFP, green fluorescent protein. PB2, polybasic region of Rac2. (B) AlphaFold3 (AF3) prediction of cytochrome b558 (NOX2 and p22phox) in complex with essential cytosolic factors Rac1Q61L–PB2, p67phox(1–212), and p47phox(1–286). (C,D) Predicted interfaces shown with contacting residues and colored by pLDDT (threshold >50). (C) Interface between the p67phox activation domain (AD) and NOX2. (D) Interface between the p22phox proline rich region (PRR) and the p47phox SH3a and SH3b domains.
In the meantime, the Python scripts are being used for the organization and analysis of the cytosolic subunits of the NOX2 complex. The ongoing work also addresses one of the known limitations of AlphaFold3, namely its limited ability to directly predict multiple conformational states, as well as the effects of point mutations and post-translational modifications.
To go beyond and complement AlphaFold3 predictions, Octavio’s approach involves increasing the number of prediction seeds in order to sample a broader conformational space. The analysis scripts under development are designed to discriminate, rank, and select representative structures suitable for subsequent molecular dynamics (MD) simulations. These tools will enable the prediction of the behavior of the NOX2 complex in specific membrane compositions, whether experimentally defined or native-like environments. This integrated modelling strategy will support the design of more targeted and informative experiments and will provide deeper insights into the dynamics and regulation of the cytosolic components of the NADPH oxidase complex.
The successful completion and full exploitation of this work will require an extension of the project duration, in order to explore the full predictive potential of the developed models and computational framework.
4. Benjamin Kusi

Mr. Benjamin Kusi is a Master’s student in Data Science and Business Intelligence at EDC Paris Business School. He has been recruited as an intern under the FEF-IA project to work on the research topic entitled “Accelerating Quantum Chemistry Calculations with Neural Networks,” for the period from 1 December 2025 to 30 April 2026.
The primary objective of Benjamin’s project is to explore machine learning–based approaches for accelerating the evaluation of two-electron (2-e) integrals in Hartree–Fock (HF) and multiconfigurational self-consistent field (MCSCF) calculations, while avoiding any increase in memory requirements. Improving the efficiency of these calculations is critical for enabling large-scale quantum chemistry simulations relevant to complex biological and chemical systems.
As an initial step, Benjamin has performed Hartree–Fock calculations on the hydrogen molecule (H₂) using a minimal Slater-type orbital (STO) basis set within the PySCF (Python-based Simulation of Chemistry Framework) software package. This foundational work establishes a reference implementation and validation framework that is essential for the next phase of the project, which involves the integration of neural network models into quantum chemistry workflows.
In parallel, Benjamin will explore the Psi4 quantum chemistry package, with the objective of modifying and extending the code to interface with and accelerate the computation of two-electron integrals. This approach will subsequently be generalized to more complex molecular systems, including large biomolecules such as proteins, where computational cost is a major limiting factor.
Given the methodological complexity and the scope of development required for this component of the project, additional time will be necessary to fully implement, validate, and optimize the proposed machine learning framework. Consequently, this work necessitates an extension of the project duration to achieve its full scientific potential.
We are preparing also for other students and researchers for expanding the scope of the project. These include:
5. Mr Samuel Kwadwo Kubiti

Using Artificial Intelligence to Estimate Hemoglobin Levels in Children from Palm Images.”
His project aims to develop a non-invasive way to detect anemia in children using photographs of the palm of the hand. Hemoglobin, which carries oxygen in the blood, is a key indicator of anemia. Today, measuring it usually requires a blood test, which can be costly and difficult to access in many parts of Ghana.
Samuel will use machine learning and image analysis to study how the color and appearance of the palm relate to hemoglobin levels. By training computer models on palm images linked to medical measurements, he aims to create a system that can estimate hemoglobin using only a smartphone photo. This could make anemia screening faster, cheaper, and easier, especially in schools and rural clinics.
Samuel is expected to be in France from 1 March to 31 May 2026. During this period, he will be supervised by Dr. Misaki Ozawa, CNRS researcher at the Laboratoire Interdisciplinaire de Physique (LIPhy), Université Grenoble Alpes, where he will receive training in artificial intelligence and data analysis.
6. Miss Ellen Acquonoo-Gyasi

Miss Ellen Acquonoo-Gyasi is a Biochemist and we are on the process of recruiting her under the FEF-IA project to work on “Comparative Study of Conventional and FLASH Radiotherapy on phagocyte Function and Pathogen Clearance: integration of AI-based Predictive Models”.
The project aims to acquire a large amount of quantitative and imaging data that will need AI-supported analysis. This project will complement Sandra’s innovative research by leveraging AI to support the interpretation and analysis of data on the specific effects of radiotherapy modalities on the immune response, a key determinant of immune defense during and after patient treatment.
7. Mr Michael Mawuli Kumah

Mr Michael Mawuli Kumah is an ICT Tutor at the Dormaa Senior High School who will embark on a scientific mission under the FEF-IA project to work on the project “Development of a robotic system to assist in sample placement at the ICP gamma source”. Michael is in-charge of the Robotic club Dormaa Senior High School in Ghana. Mr Michael is expected to arrive In France from 1 March 2026- 30 April 2026.
